Introduction
It is now well known that root colonizing bacteria, which have properties of N2 fixation, production of phytohormones, enhancement of nutrient uptake or suppression of pathogens etc., are included into plant growth-promoting rhizobacteria (PGPR) defined by Kloepper and Schroth (1978). Much potential benefits of these soil microorganisms colonizing plant root have been extensively evaluated in low-input agriculture because they enable partial replacement of chemical fertilizers and other chemicals (Baldani et al., 2001; Dashti et al., 1998; de Freitas and Germida, 1990; Kloepper and Schroth, 1981; Perez-Montano et al., 2014). Especially in Asian rice agriculture, the role of PGPR is attracting attention not only of agricultural scientists but also of rice farmers.
The best known microorganisms with PGPR activity are the bacteria belonging to the genera Azospirrilum (Okon, 1985), Pseudomonas (Kloepper et al., 1980) and Bacillus (Backman et al., 1994; Park et al., 2015). In most agroecosystems, these bacteria exert their role in plant growth promotion either directly or indirectly. Their indirect facilitation of plant growth is attributed to a plethora of mechanisms, such as by enabling the growth of other beneficial soil microorganisms or by preventing phytopathogens from inhibiting plant growth (Lugtenberg and Kamilova, 2009; Shen, 1997; Vessey, 2003). The direct action may be specified by N2 fixation, phytohormones, phosphate solublization etc. (Lavakush et al., 2014; Lugtenberg and Kamilova, 2009; Vessey, 2003). Furthermore, the PGPR like Azospirillum, Sinorhizobium meliloti, etc., possibly affect root respiration, which in turn leads to increase in root growth (Phillips et al., 1999; Vedder-Weiss et al., 1999). This positive effect by PGPR on root growth provides the host plants with a high nutrient uptake potential, resulting in overall growth enhancement and higher yields (Barber and Silverbush, 1984; Lavakush et al., 2014; Perez-Montano et al., 2014). Ever since Hiltner (1904) observed that living roots stimulate soil microorganisms, varieties of PGPR have been successfully employed in agricultural practices to enhance the production of potato (Kloepper et al., 1980), soybean (Dashti et al., 1998), wheat (de Freitas and Germida, 1990), sugar beets (Suslow and Schrith, 1982), rice (Baldani et al., 2001) etc..
Because of the beneficial effects of PGPR on plant growth, practical studies consistently stress the importance of the root colonization by PGPR for their optimal performance in a wide range of host plants and edaphic conditions (Kloepper and Beauchamp, 1992; Shen, 1997; Vessey, 2003). Root colonization by the introduced bacteria is a crucial step to enable the establishment of their association with the host plant in the presence of predominant indigenous microbial community already present in the rhizosphere (Hiltner, 1904). In this respect, the concept of quorum in the initial bacterial inoculum to meet the need for a threshold level imply the importance of root colonization for the expectation of positive yield response (Persello-Cartieaux et al., 2003). Of course, the contributions of root colonizing PGPR to plant growth appear to be diverse depending on their colonizing patterns, which can be categorized into their rhizospheric and endophytic relationship with host plant. The former PGPR colonizing state has an absolute advantage for root growth, the availability of plant nutrients and protection of roots from plant pathogens in soils, whereas the latter state is a desirable trait for directly providing the host plant with nitrogen, etc. (Baldani et al., 1996; Muthukumarasamy et al., 2003). Thus the use of a PGPR preparation with either a single or mixed population of strains having an ability to colonize surface as well as interior of the plant is recommended for stimulating plant growth through various synergistic and potential beneficial effects of PGPRs (Kozhevin and Korchmaru, 1995; Muthukumarasamy et al., 2003). Especially in the case of rice, incorporation of the endophytic diazotrophs such as Herbaspirillum seropedicae and Burkolderia brasilensis etc., which are capable of inhabiting the stem and leaves of plant, into PGPR inoculum can augment nitrogen to the host plant in temperate as well as tropical regions (Baldani et al., 2001). Despite some successes with the inoculation of plant growth-promoting bacteria, there are still several problems associated with soil texture (Mahaffee, 1991), soil nitrogen (Caballero-Mellado et al., 1995; Muthukumarasamy et al., 2002), soil water (Mayak et al., 2004; Timmusk and Wagner, 1999), and plant growth stage (Ladha et al.,1988; van Nieuwenhove et al., 2000; Sethunathan et al., 1983) need to be solved for the establishment of successful association between host plant and PGPR.
The purpose of this study was to identify the promising rhizobacteria with a property of stimulating the growth of rice in order to develop a biofertilizer for reducing the dependency on chemical fertilizer, and enabling an environmentally friendly agriculture. During the investigations a number of potential rhizobacteria were screened using a bioassay for plant growth promotion and N2-fixing activity, etc. Subsequently the root colonization, and yield response of a Korean rice variety to inoculation with selected isolates were determined under pot and field conditions.
Materials and Methods
Isolation of rice rhizobacteria Twenty Korean soils and five Russian soils were used for isolation of plant root colonizing bacteria (Table 1). For isolation of rice rhizobacteria, hulled rice seeds of the Japonica variety Goamibyeo were sterilized with 3% H2O2 for 10 minutes, and germinated on 0.6% water agarose plates for 24 hours at 28°C. Subsequently five-gram of a soil sample was placed at a distance of 1-2 cm parallel to the roots of rice seedlings. Thus soil microorganisms were allowed to move towards the nutrient source of root exudates for 24 hours. Then the rice roots were crushed and streaked with a sterile loop on potato dextrose agar (PDA) plates for isolating colonies.
Identification of rice rhizobacteria The putative PGPR strains were identified by the comparative analysis of 16S ribosomal RNA gene sequence. For the analysis, chromosomal DNAs were isolated by a versatile quick-prep method with some modification (Pospiech and Neumann, 1995). Cells (about 0.7 ml) grown in TSB were centrifuged, rinsed with TE and suspended in 0.4 ml of SET buffer (75 mM NaCl, 25 mM EDTA, 20 mM Tris, pH 7.5). Three volumes of 5 M NaCl and 1 volume of chloroform were added and incubated at room temperature for 30 min and then centrifuged at 4500 g for 15 min after which the aqueous phase was collected. The chromosomal DNA was precipitated by addition of 1 volume of iso-propanol and gentle inversion. DNA was transferred, rinsed with 70% ethanol, dried under vacuum, and dissolved in a suitable volume of distilled water and then treated with 20 µg per ml of RNase at 37°C for 1 hr. The samples were extracted with phenol:chloroform:isoamyl alcohol (25:24:1), precipitated with 2.5 volumes of cold ethanol and 1/10 volumes of 3M sodium acetate. The pellets were washed with 70% ethanol, dried and dissolved in TE buffer. The fragments of each 16S rRNA genes were amplified using oligonucleotide primers complementary to highly conserved regions of the bacterial 16S rRNA genes. The forward primer was 5'-AGAGTTTGATCCTGGCTCAG-3' (positions 8-27, according to the Escherichia coli numbering system), the reverse primer 5'-AAGGAGGTGATCCAGCCGCA-3' (positions 1541-1522). PCR products were purified using a QIA quick PCR Purification Kit (Qiagen), according to the manufacturer’s instructions. Purified PCR products were partially sequenced by using the ABI Prism Big Dye Terminator Cycle Sequencing Ready Reaction Kit and an Applied Biosystems 377 DNA sequencer, using the protocols of the manufacturer (Perkin-Elmer). The sequencing primers used were *Gamma, Gamma and PD (Coenye et al., 1999). Sequence assembly was performed using the program AutoAssembler (Applied Biosystems).
Physiolgical characterization of isolates Bioassay test for plant growth promotion by rhizobacterial strain : The bioassay was conducted on maize seedlings. Maize seeds were washed with tap water for two hours, placed in a plastic tray with a moist filter paper, and allowed to germinate in dark at 28°C in an incubator. After 48 hours of growth the tips of the maize roots were dipped into bacterial suspension for 1.5 hours. In the control, the roots were treated with sterile growth medium. Subsequently the maize seedlings were placed on a moist filter paper in a tray, with all root tips aligned to a starting line, covered with another filter paper and incubated at 28°C. After 24 hours the roots, which crossed the starting line were cut off and their length was measured. Ten maize seedlings were used for testing growth promotion ability of one strain of rhizobacterium.
Acetylene reduction assay : N2-fixing ability of rhizobacteria was analyzed using acetylene reduction technique (Kang et al., 1991). For measuring acetylene reducing activity, the rhizobacterial strains were inoculated in to the bottles containing LGI medium and covered with caps of serum stoppers. The inoculated bottles were incubated at 28°C and used in time course experiments. Acetylene was injected into 4 bottles per each strain and the bottles were incubated for 30 minutes at 28°C. Subsequently the gas phase from the bottles was sampled to determine ethylene production by gas chromatography (Shimadzu model GC-17A).
Bioassay for determining antimicrobial activity of rhizobacteria : Biocontrol activity of rhizobacterial isolates was assessed on fungal and bacterial phytopathogens. Conidial (5 ml of 1×106 conidia ml-1) and bacterial (5ml of 1×108 cfu ml-1) suspensions of plant pathogenic fungal and bacterial strains, respectively, were separately added to 500 ml flasks containing 250 ml Czapek agar at 45°C, carefully mixed and poured into petri dishes (25 ml each). Separately, rhizobacterial inoculum was prepared from a stationary bacterial culture grown on liquid potato dextrose medium for 5 days at 28°C without shaking. Inhibition of fungal and bacterial growth by rhizobacteria was assessed by placing 100 µl inoculum into 8 mm wells cut into Czapek agar plates having fungal or bacterial phytopathogens. Antifungal and antibacterial activities were defined by the diameter of the fungal and bacterial growth inhibition zone around wells observed after an incubation for 2-4 days at 28°C.
Phosphate solubilization activity : In order to measure phosphate solubilization activity, rhizobacterial strains were inoculated onto agar medium containing glucose, 10.0 g, yeast extract, 0.50 g, calcium chloride, 0.10 g; hydroxyapatite, 4.0 g, agar, 15.0 g per liter of distilled water at pH 7.0, and incubated for 4 days at 28°C. Formation of transperant zones around the rhizobacterial colonies indicated the phosphate solublizing activity of the bacteria.
Protease and pectinase activities : Rhizobacterial isolates were inoculated onto LB or Potato agar plates supplemented with 4% milk and incubated for 3 days. The transparent zones developed around the bacterial colonies indicated the protease activity of the rhizobacterial strans. Pectinase activity was estimated by inoculating of rhizobacterial isolates onto LB supplemented with 2% pectin. The turbid zones formed around the bacterial colony after 5 days of incubation at 28°C indicated pectinase activity.
Bacterial motility : To evaluate bacterial motility a plate with LB 0.3% agar swarming was inoculated using a toothpick with rhizobacterial isolate cultured on LB 0.8% agar plate for two days. The motility was quantified by measuring the semidiameter of the bacterial colony.
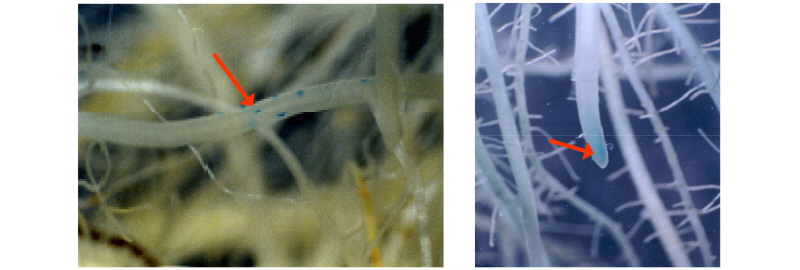
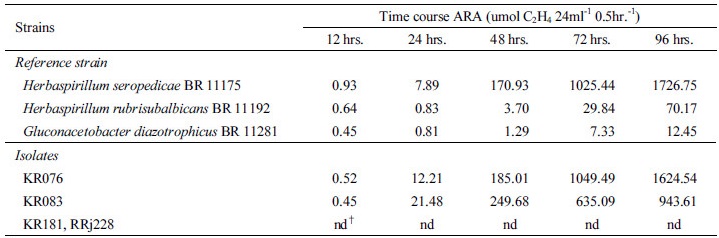
Rice root colonization by the rhizobacterial isolates Ability of the rhizobacterial strains to colonize the roots was evaluated on several rice varieties belonging to Japonica and Korean Tongil varieties grown in pot. Root colonization was investigated adopting two types of inoculation procedures: 1. Inoculation with rhizobacteria at a density of 2.2×104 cfu ml-1 of paddy water three days after transplantation of 30 days old rice seedlings; 2. Inoculation by soaking the rice roots in rhizobacterial suspension of 1.1×105 cfu ml-1 for 6 and 12 hours at room temperature. The rhizobacterial strains KR076, KR083, and RRj228 were cultured in liquid Potato Dextrose medium, while the strain KR181 was grown in YMA medium. For differentiation of rhizobacterial strains from native soil microorganisms in rice roots, rhizobacteria were tagged with gusA by conjugative mobilization of a plasmid construct, mTn5SSgusA20 (Kang et al., 1997). Both rhizoplane and endophytic colonization of roots were assessed in the plants inoculated by the soaking method, while only the endophytic colonization was evaluated in the plants inoculated in paddy water. Endophytic colonization was observed under stereoscopic microscope after root staining with GUS assay solution (Kang et al., 1997). In order to evaluate the endophytic and rhizoplane colonization by the inoculated rhizobacteria, root samples were ground, diluted and plated onto PDA medium (per 1 liter) containing 1 ml of 2% X-gluc (5-bromo-4-chloro-3-indolyl-D-glucuronic acid), and gus-expressing rhizobacteria were enumerated.
Rice yield response Experiments on rice yield response to rhizobacterial inoculation were conducted under pot and field conditions. The rhizobacterial strains KR076, KR083, and RRj228 grown on PDA medium and the strain KR181 grown on YMA medium were used as inoculants. In pot trials, the rice variety Daesanbyeo was tested with single strain inoculation, while Mananbyeo was evaluated in coinoculation studies with RRj228 and KR076 or KR083. Inoculation was performed by dipping the roots of 30 days old rice seedlings in bacterial suspension of 2.0×106 cfu ml-1 for 12 hours. The inoculated seedlings were then transplanted to pots containing soil. Uninoculated plants treated with sterile medium before transplantation were served as control. Yield response of inoculated rice plants was determined under fertilization regime of N, P2O5 and K2O at a half dose level, and compared it with yield response of uninoculated plants fertilized with a standard full dose of NPK at a level of N 110, P2O5 45, and K2O 57 kg ha-1. In field trials, growth response of the rice variety Junambyeo was evaluated using single strain inoculation with RRj228 at a 70% dose level of nitrogen, as well as by coinoculation with KR083/KR181 or KR083/RRj228 at a half dose level of nitrogen. In the former case, roots of the rice seedlings were dipped in a bacterial suspension of 1.8×106 cfu ml-1 for 12 hours, and then grown in nursery box for 30 days before transplanting them into the field. In the latter case, 30 day-old rice seedlings were treated with inoculum containing 1.0×106 cfu ml-1 like the former. Inoculated rice plants were transplanted to paddy soils of a fine silty loamy textured alluvial deposit, and the yield responses to inoculation were evaluated.
Results
Isolation, selection, and identification of rice rhizobacteria A total of 172 rhizobacteria were isolated from Korean and Russian soils. The isolates obtained from Korean rice soils were given a prefix KR, while those isolated from Russian rice soils were provided with a prefix RR (Table 1).
Among these isolates, KR076, KR083, KR181 from Korean soils, and RRj228 from Russian soils were found to be potential PGPR strains. Bioassays on maize seedlings showed that the strains KR076, KR083, and KR181 increased the root length by 21.3%, 26.3%, and 15.0%, respectively, when the plants were inoculated with 50-fold diluted cell suspensions of stationary phase cultures controls as compared to uninoculated one (Table 2).
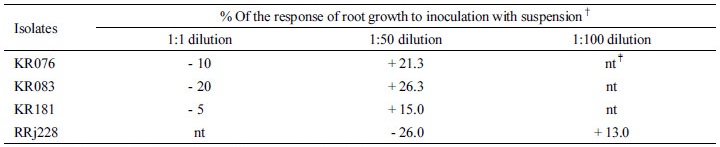
†Determined 1 day after inoculation on filter paper inside plastic trays.
‡Not tried.
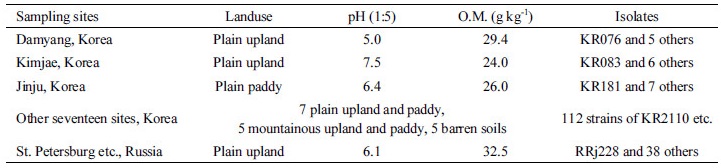
On the contrary, RRj228 inhibited the root growth by 26% when inoculated with 50-fold diluted cell suspension, but stimulated the growth by 13% when inoculated with 100-fold diluted cell suspension. With regard to N2-fixing ability, the strains KR076 and KR083 showed higher acetylene reducing activity (ARA) than the best reference diazotroph Herbaspirillum seropedicae BR11175 (Table 3). In cultures incubated for 48 hours at 28°C, KR076 and KR083 strains exhibited more than 50-fold higher activities than Herbaspirillum rubrisubalbican BR11192 and Gluconacetobacter diazotrophicus BR11281, and 1.1 - 1.5-fold higher than Herbaspirillum seropedicae BR11175. All four above mentioned isolates were found to be effective phosphate solublizers.
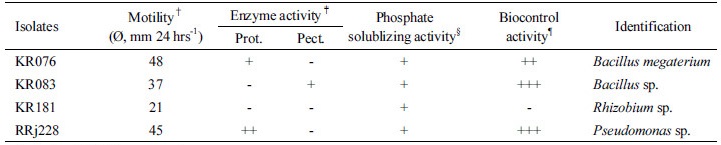
†Tested on 0.3% Agarose media.
‡Prot, protease; Lip, lipase; Pect, pectinase.
§Phosphate solublizing activity was evaluated as a clear zone on a agar medium with hydroxyapatite.
¶Biocontrol activity was estimated as inhibition zones of isolates against plant pathogen Rhizoctonia solani, Fusarium culmorum, Pythium ultimum, Erwinia carotovora, Pseudomonas syringiae etc.. *Inhibition zone +++, >15 mm; ++, 10 mm
In bioassays on biocontrol activities, the strains KR076, KR083, and RRj228 showed positive reactions. In contrast, the strain KR181 did not show any effect on the plant pathogens Erwinia carotovora, Fusarium culmorum, Pythium ultimum, Pseudomonas syringiae, Rhizoctonia solani etc. Among the efficient biocontrol isolates, RRj228 was found to be the most effective on the plant pathogens tested. Assessment of hydrolytic enzymatic activities, which mediate bacterial invasion of plant tissues, revealed that the strains KR076 and RRj228 produced protease, while the isolate KR083 produced pectinase. On the contrary, the strain KR181 did not produce either enzyme. In motility test for root colonization, the strain KR076 showed higher mobility than other three isolates. KR076 exhibited a mobility of 24 mm in 24 hours in agarose medium, while RRj228, KR083 and KR181 could cover only a distance of 22.5, 18.5 and 10.5 mm, respectively, within 24 hrs.
Based on the comparative analysis of 16S rRNA gene sequences, the strains KR076, KR083, KR181 and RRj228 were identified as Bacillus megaterium, Bacillus sp., Rhizobium sp., and Pseudomonas sp., respectively; thus these strains were designated as Bacillus megaterium KR076, Bacillus sp. KR083, Rhizobium sp. KR181, and Pseudomonas sp. RRj228.
Rice root colonization Root colonization by the rhizobacteria KR076, KR083, KR181 and RRj228 was assessed in both Japonica and Korean Tongil type rice varieties (Table 5). In both groups of rice varieties, no endophytic colonization was observed till 10 days after inoculation. However, Japonica varieties showed internal colonization 20 days after inoculation; the rhizobacterial strains KR076, KR083 and KR181 were able to colonize four Japonica rice varieties, namely Goamibyeo, Gumobyeo, Hwaseongbyeo and Junambyeo, whereas the strain RRj228 colonized all five Japonica varieties including Ilpumbyeo. On the other hand, none of these rhizobacterial strains were able to colonize Korean Tongil type of rice varieties such as Andabyeo, Arumbyeo, Dasanbyeo and Samgangbyeo. In the case of coinoculation of Japonica rice with KR083/KR181 or KR083/RRj228, number of varieties with internal colonization of rice roots was diminished in both Japonica and Korean Tongil type compared with the tendency exhibited by the single inoculant. When the rice roots were inoculated by soaking into rhizobacterial suspension for six and twelve hours, however, there was not big difference between Japonica and Korean Tongil type rice varieties in population living on rhizoplane and/or inside roots as in Table 6. Average population density of the rhizobacterial strains colonizing root surface was also more in Japonica than Korean Tongil type rice varieties. This tendency of population density was more obvious in the rice roots dipped in rhizobacterial suspension for 12 hours than for 6 hours. In case of the rice roots treated for 12 hours, average population density of rhizobacteria was 53.5×103 cfu g․fw-1 for Japonica and 37.9×103 cfu g․fw-1 for Korean Tongil type. On the other hand, in the roots treated for 6 hours, the average number was only 31.8×103 cfu g․fw-1 in Japonica and 20.6×103 cfu g․fw-1 in Korean Tongil type varieties.
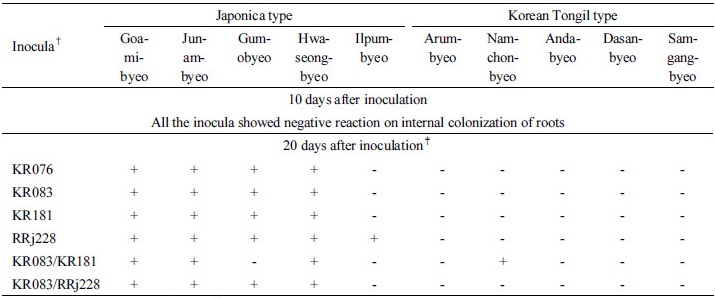

†Inoculum size was 2.2 × 104 cfu ml-1in paddy water.
‡+ and - mean root colonization and no colonization, respectively.
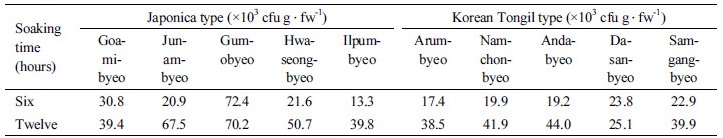
†Inoculum and its size were strains KR083/RRj228 and 1.1 × 105 cfu ml-1, respectively.
Rice yields In pot trials, at half a dose of N, P2O5, and K2O fertilizer, Daesnbyeo showed positive yield response with a single strain inoculation with KR181 or RRj228, while it exhibited a negative response with the strains KR076 or KR083 (Table 7). The yield response of Daesanbyeo was 35.1% (p<0.01) and 33.0% (p<0.01) higher when inoculated with KR181 and RRj228, respectively, as compared to control. Moreover, inoculation with the strain KR181 promoted yields equivalent to that obtained with a half dose of N, P2O5, and K2O fertilizer. In Mananbyeo rice variety, when compared to Daesanbyeo, KR083 and KR076 produced positive effects by the coinoculation together with RRj228 on yield response. KR083/RRj228 combination of coinoculation promoted a higher yield response of 14.9% (p<0.05) as compared to the combination KR076/RRj228, which enhanced yield only by 7.1% (p<0.05).
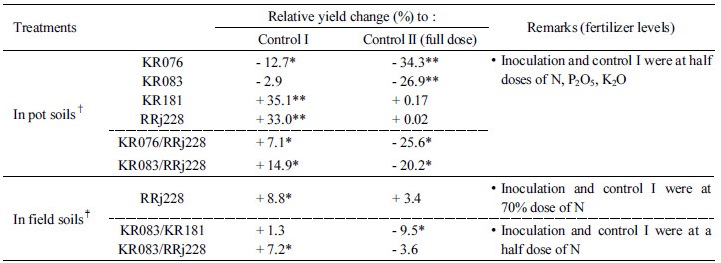
†Rice variety was Daesanbyeo for KR076, KR083, KR181 and RRj228; Mananbyeo for KR076/RRj228 and KR083/RRj228.
‡Rice variety was Junambyeo for all inocula.
* and ** mean least significant difference from control plot at 1% and 5% levels by DMRT, respectively.
In field trials, upon inoculation with RRj228, rice yields increased by about 8.8% (p<0.05) and 3.4% when supplemented the crop with 70% dose and a full dose of nitrogen fertilizer, respectively. In the rice variety Junambyeo, at a half dose of nitrogen fertilizer, coinoculation with KR083/RRj228 enhanced yield by about 7.2% (p<0.05), while it increased by 1.3% with KR083/KR181 combination. The yield from the coinoculation at a half dose of N, however, was 3.6% and 9.5% (p<0.05) less than those from uninoculated control at a full dose of N when the inocula were KR083/RRj228 and KR083/KR181, respectively.
Discussion
Ever since the associative relationship between plant roots and soil microorganisms was observed by Hiltner (1904), concerted attempts were made by several researchers to isolate microorganisms useful for agriculture, and some of them were developed as biofertilizers (Bashan et al., 2014). In this study, Bacillus megaterium KR076, Bacillus sp. KR083, Rhizobium sp. KR181, and Pseudomonas sp. RRj228 were identified as plant growth- promoting rhizobacteria (PGPR) based on in vitro evaluation of their beneficial effects, such as plant growth- promotion, N2-fixation, P solublization, biocontrol, etc.. Most important plant growth-promoting activity of rhizobacterial strains is related to root growth (Perez-Montano et al., 2014; Persello-Cartieaux et al., 2003). In general, plant growth-promotion by PGPR is highly correlated with the increase of root surface area (Bertrand et al., 2000; Okon and Vanderleyden, 1997; Pacovsky, 1990). As a consequence of phytohormones such as indol-3-acetic acid and cytokinins produced by PGPR, root proliferation including branching of roots is increased, which enable enhanced uptake of a variety of nutrients (Barazani and Friedman, 1999; Persello-Cartieaux et al., 2003; Timmusk et al., 1999). In the present study the observed increase in root growth in maize (Table 2) is likely due to phytohormones production by rhizobacteria. Persello-Cartieaux et al. (2003) showed that inoculation with increasing doses of Pseudomonas strains has a positive impact on root growth up to certain concentration of bacteria, and beyond which the bacterial inoculation led to deleterious effects. Among four strains tested in the present study, Pseudomonas strain RRj228, which promoted good root growth at the lowest cell density seem to have a great potential for rice farming. In fact, inoculation of RRj228 with a concentration of 105 cfu ml-1 resulted in higher shoot dry weight in rice than when inoculated with 107 cfu ml-1 (Kang, Unpublished data). Dobbelaere et al. (2002) also reported that the inoculum at a concentration of 105 - 106 cfu plant-1 of Azospirillum brasilense Sp245 produced a positive effect on wheat as compared to the inoculum strength of 107 - 108 cfu plant-1.
In order for PGPR to promote growth of the plant, root colonization is emphasized as a primary step for the initiation of associative relationship between microorganism and its host plant (Hiltner, 1904; Kloepper and Beauchamp, 1992; Lugtenberg et al., 2001; Lugtenberg and Kamilova, 2009; Perez-Montano et al., 2014). The fact that rhizobacterial strains of the present study are able to colonize the roots of Japonica rice varieties rather than Korean Tongil type (Table 5 and Fig. 1) indicates that rhizobacteria are able to associate only with compatible host plant varieties. In spite of better association, no rhizobacteria were observed in apoplastic spaces (Kovtunovych et al., 1999; Verma et al., 2001) even in Japonica varieties. McCulley (2001) reported that heavily suberized and/or lignified cell layers in certain plants could form a barrier to the bacteria. Rhizobacterial strains KR076 and RRj228 of the present study showed protease activity, while the strain KR083 possessed pectinase activity. On the contrary, the strain KR181 didn’t show either protease or pectinase activity (Table 4).
However, the abilities of rhizobacterial strains for endophytic colonization in rice roots seem to be decline when coinoculated with KR083/KR181 or KR083/RRj228 compared to single inoculation (Table 5).
When the colonization of rhizoplane and/or inside roots was determined, almost biased pattern of inside root colonization to Japonica in not only paddy water inoculation (Table 5) but also root inoculation trial (Kang, Unpublished data) was largely changed although population density averaged in whole roots of Korean Tongil type was as much as 65 and 71% of that in Japonica with the soaking inoculation for 6 and 12 hours, respectively (Table 6). The fact that population density in roots was still high in Japonica indicated that the gap of value originated from the cells living inside roots of Japonica rice plant. The root colonizing PGPR, particularly Bacillus and Pseudomonas sp., were found to suppress various plant diseases caused by soil-born pathogenic microorganisms (Handelsman and Stabb, 1996; Lugtenberg and Kamilova, 2009). Likewise, Bacillus sp. strain KR083 and Pseudomonas sp. strain RRj228 were found to be effective root colonizers with biocontrol activities
Results of the present study demonstrated that yield responses of rice to inoculation with rhizobacteria are dependent on rhizobacterial inoculant composed of single or mixed strains (Table 7). Yield responses to inoculation of rhizobacteria also depend on rice cultivars and fertilizer levels. These facts corroborate the findings of Dashti et al. (1998), Muthukumarasamy et al. (2002) and van Nieuwenhove et al. (2000). Rice yield was positively affected by inoculation with Rhizobium strain KR181 and Pseudomonas strain RRj228, which have no detectible N2-fixation activity, but have strong plant growth promotion activity at low population concentrations. Furthermore, the strain RRj228 possessed phosphate solublization activity (Kang, Unpublished data). On the other hand, rice yield was negatively affected by Bacillus strains KR076 and KR083 although they exhibited high N2-fixing activities in free living state and colonized rice roots endophytically (Table 5). As a result, the yield response of rice to coinoculation revealed dependence on a component strain; for example, decreased positive effect by superior strains such as KR181 or RRj228 when coinoculated with inferior strains like KR076 or KR083. Despite of much research to verify that growth promotion of rice plant by PGPR is due to their ability to fix N2 in situ, which include Azospirillum sp. (Malik et al., 1997), Azoarcus sp. (Egener et al., 1999), Burkholderia sp. (Baldani et al., 2001), and Herbaspirillum sp. (James et al., 2002), vast data points to the fact that growth promoting effects of PGPR originate predominantly from other mechanisms such as phytohormon production, effects on root morphology, etc. (Bertrand et al., 2000; Lavakush et al., 2014; Lugtenberg and Kamilova, 2009; Pacovsky, 1990; Persello-Cartieaux et al., 2003). It still remained obscure that the lack of nitrogenase activity in the rice roots occured despite the successful colonization by N2-fixing rhizobacteria. According to van Dommelen et al. (1997), no significant contribution of N2-fixation by PGPR to host plant is thought to be involved in the reabsorption of NH4+ released as a consequence of NH3 diffusion through the bacterial membrane. In current study, the fact that Bacillus strains negatively contributed to yield of paddy rice even though they harbored the property of both phytohormon production like IAA (Kang, Unpublished data) and N2-fixing activity supposedly indicated that there was an influence of unique environment of submerged paddy soils (Bashan, 1999; van Nieuwenhove et al., 2000). As a result, KR076 and KR083 strains produced a positive yield responses for crops, such as barley, potato, perilla, cucumber, etc., grown in upland soils which generally contain the water of field capacity (Kang, Unpublished data).
In conclusion, to expect the consistent beneficial effects of PGPR contributing to sustainable rice agriculture, the improvement of limiting factors using the diversity of the microbial community is required in terms of the influence of host cultivars, climatic and edaphic conditions (Kozhevin and Korchmaru, 1995; Muthukumarasamy et al., 1999; Nadeem et al., 2014; Shen, 1997).